Ciliome Gene Expression Reference
Welcome!
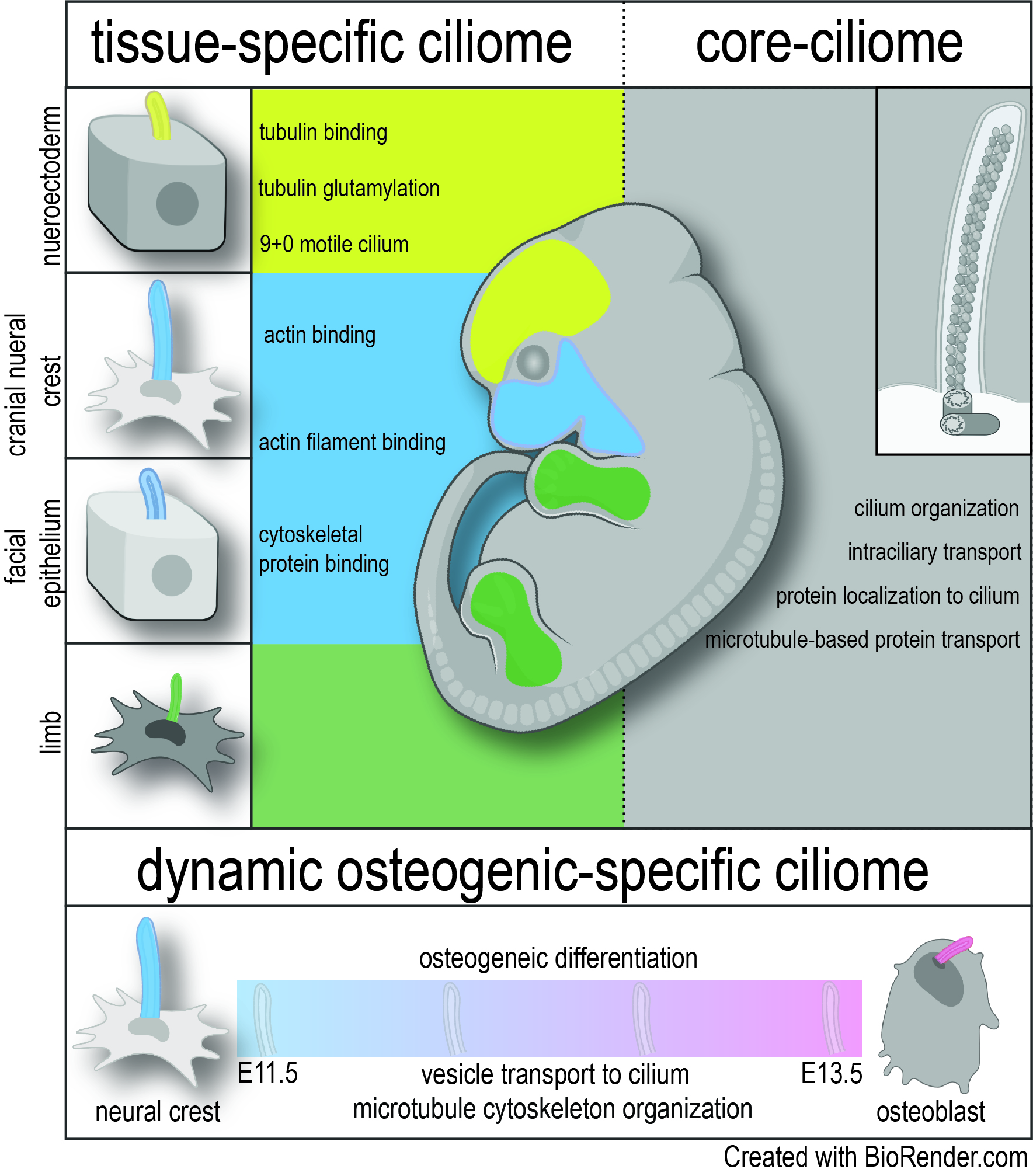
Primary cilia are nearly ubiquitous organelles that transduce molecular and mechanical signals. While the basic structure of the cilium and the cadre of genes that contribute to ciliary formation and function (the ciliome) are believed to be evolutionarily conserved, the presentation of ciliopathies with narrow, tissue-specific phenotypes and specific molecular readouts suggests an unappreciated heterogeneity within this organelle.
Herein, we consolidated three established ciliary databases: The Syscilia Gold Standard, CiliaCarta, and Cildb [1,2] with two ciliogenesis modulator screens [3, 4] to generate a curated ciliome. Expression of genes that make up the ciliome was then profiled across various embryonic tissues and time points to test the hypothesis that the ciliome is heterogenous throughout development.
Here, we provide access to our transcriptomics resources detailing tissue-specific ciliome heterogeneity in an easily searchable database.
- Arnaiz, O., et al., Remodeling Cildb, a popular database for cilia and links for ciliopathies. Cilia, 2014. 3: p. 9.
- Arnaiz, O., et al., Cildb: a knowledgebase for centrosomes and cilia. Database (Oxford), 2009. 2009: p. bap022.
- Kim, J., et al., Functional genomic screen for modulators of ciliogenesis and cilium length. Nature, 2010. 464(7291): p. 1048-51.
- Wheway, G., et al., An siRNA-based functional genomics screen for the identification of regulators of ciliogenesis and ciliopathy genes. Nat Cell Biol, 2015. 17(8): p. 1074-1087.
Contact us:
Samantha Brugmann, Samantha.Brugmann@cchmc.org
Kevin Peterson, kevin.peterson@jax.org
Developed by Konrad Thorner
For technical issues please see the GitHub page
Database Instructions
- Some columns are hidden by default. Additional columns can be viewed using the 'Column Visibility' dropdown.
- Results can be saved using the CSV and Excel buttons for further analysis.
- View genes of interest by entering one or more comma-separated genes in the 'Gene Search' box.
- For more advanced searches, use the filters at the top of each column. Regular expressions can also be applied.
- Search for multiple entries by separating with ' | '. Ex. Foxj1|Dnah9
- Search for entries that start with a pattern using ' ^ '. Ex. ^A returns all entries starting with A
Database Legend
- human_ensembl_id : Human Ensembl gene identifier
- human_gene_symbol : Human gene symbol
- human_synonym : Human gene synonym
- CilDB : CilDB-X denotes established ciliopathy genes, Y denotes candidate ciliopathy genes
- SysCilia : SysCilia-X denotes confirmed, Y denotes potential ciliary gene
- ciliaCarta_rank : CiliaCarta Rank
- CiliaCarta_score : CiliaCarta Score
- ciliogenesis : Ciliogensis-denotes validated positive or negative regulators published by Kim et al., 2010
- siRNA : siRNA-X denotes validated ciliary gene published by Wheway et al., 2015
- cilia_motility_class : Cilia motility classificaiton based upon Partir et al., 2020
- mgi_accession_id : Mouse MGI accession identifier (http://www.informatics.jax.org/)
- mouse_symbol : Mouse gene symbol
- mouse_synonym : Mouse gene synonym
- mouse_name : Mouse gene name
- differential_expression_group : Gene sets defined as differentially expressed in pairwise comparison (>2-fold change and FDR < 0.05) between any two tissues (DE_CILIOME) or not differentially expressed (non-DE), see Fig.1A
- tissue_specificity : Tissue specificity of differentially expressed genes, see Fig. 2B
- facial_prominence : Facial prominence specificity of differentially expressed genes, see Fig. 2D
- cell_type_specificity : Cell type specific expression of DE_ciliome genes from scRNA-seq profiling in E11.5 mandibles, see Fig.3A
- osteo_group : Genes showing increased gene expression from E11.5 to E13.5 mandibles correlating with onset of osteogenesis
- neural_specificity : Neural tube specificity, see Supplemental Fig. 2
- mp_id_impc : Mammalian Phenotype (MP) identifiers for statistically significant hits in mutant mice characterized by the International Phenotyping Mouse Consortium
- mp_term_impc : Mammalian Phenotype (MP) terms for statistically significant hits in mutant mice characterized by the International Phenotyping Mouse Consortium
- impc_link : Link to IMPC generated data
- oe_lof_upper : The observed to expected loss-of-function variants (LOEUF), according to gnomAD
- oe_lof_upper_bin : The deciles for the scores in oe_lof_upper
- Ishikawa et al. 2011 : Proteins identified by Ishikawa et al. 2011 [1]
- Mick et al. 2015 : Proteins identified by Mick et al. 2015 [2]
- May et al. 2021 : Proteins identified by May et al. 2021 [3]
- Ishikawa, H., et al., Proteomic analysis of mammalian primary cilia. Curr Biol, 2012. 22(5): p. 414-9.
- Mick, D.U., et al., Proteomics of Primary Cilia by Proximity Labeling. Dev Cell, 2015. 35(4): p. 497-512.
- May, E.A., et al., Time-resolved proteomics profiling of the ciliary Hedgehog response. J Cell Biol, 2021. 220(5).