SLICE
Determining Cell Differentiation and Lineage based on Single Cell Entropy
The SLICE (Single Cell Lineage Inference Using Cell Expression Similarity and Entropy) algorithm consists of two major steps: (1) measuring cell differentiation states based on the calculation of single cell entropy (scEntropy) and (2) predicting cell differentiation trajectories by ordering single cells according to their scEntropy-derived differentiation states.
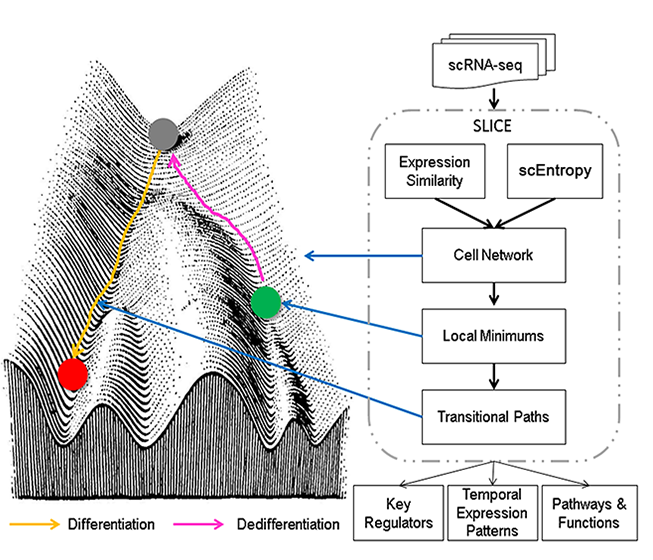
SLICE hypothesizes that entropy inversely correlates with cell differentiation state: high entropy is associated with higher functional uncertainty and more potential (cell stemness), while low entropy is associated with differentiated states with well-defined cell fates and more homogeneous functions. Therefore, scEntropy quantifies the differentiation state of a given cell by measuring the uncertainty in the activation of its cellular functions.
Using the differentiation states of individual cells measured by scEntropy, SLICE identifies relative stable cell states, defined as the centroids of the cells with local minimum entropies, and then predicts transitional paths following entropy reduction between stable cell states to reconstruct cell lineages.
• Minzhe Guo*, Erik L. Bao*, Michael Wagner, Jeffrey A. Whitsett, Yan Xu. 2016. SLICE: determing cell differentiation and lineage based on single cell entropy. Nucleic Acids Research. doi:10.1093/nar/gkw1278. (* co-first author)