LGEA News
New web development
- LAM Cell Atlas has several shiny applications for network analysis, Spatial Transcriptomics, and Single-Cell Transcriptomics under development.
- The project page for our new paper with samples from ACDMPV and control patients is available online.
- New data is available in LAM Cell Atlas Web-Portal (LCA) including integrative scRNA kidney samples.
- Check out LAM Cell Atlas Web-Portal (LCA) for LAM projects and datasets with lung, kidney and uterus cells from LAM patients.
- Check out LGEA (LungMAP phase 3) for more datasets from normal and disease lungs.
- LGEA provides query links to access DESCARTES (Human Gene Expression During Development) website in LungDTC panel.
- Hua He, Sheila M Bell, Ashley Kuenzi Davis, Shuyang Zhao, Anusha Sridharan, Cheng-Lun Na, Minzhe Guo, Yan Xu, John Snowball, Daniel T Swarr, William J Zacharias, Jeffrey A Whitsett PRDM3/16 regulate chromatin accessibility required for NKX2-1 mediated alveolar epithelial differentiation and function. Nature Communications 15, 8112 2024. Nature Communications
- Nathan Gaddis, Joshua Fortriede, Minzhe Guo, Eric E. Bardes, Michal Kouril, Scott Tabar, Kevin Burns, Maryanne E. Ardini-Poleske, Stephanie Loos, Daniel Schnell, Kang Jin, Balaji Iyer, Yina Du, Bing-Xing Huo, Anukana Bhattacharjee, Jeff Korte, Ruchi Munshi, Victoria Smith, Andrew Herbst, Joseph A. Kitzmiller, Geremy C. Clair, James P. Carson, Joshua Adkins, Edward E. Morrisey, Gloria S. Pryhuber, Ravi Misra, Jeffrey A. Whitsett, Xin Sun, Trevor Heathorn, Benedict Paten, V. B. Surya Prasath, Yan Xu, Tim Tickle, Bruce J. Aronow, Nathan Salomonis LungMAP Portal Ecosystem: Systems-level Exploration of the Lung. Am J Respir Cell Mol Biol 70, 129-139. ATS Journals
- Soumyaroop Bhattacharya, Jacquelyn A. Myers, Cameron Baker, Minzhe Guo, Soula Danopoulos, Jason R. Myers, Gautam Bandyopadhyay, Stephen T. Romas, Heidie L. Huyck, Ravi S. Misra, Jennifer Dutra, Jeanne Holden-Wiltse, Andrew N. McDavid, John M. Ashton, Denise Al Alam, S. Steven Potter, Jeffrey A. Whitsett, Yan Xu, Gloria S. Pryhuber, Thomas J. Mariani Single-Cell Transcriptomic Profiling Identifies Molecular Phenotypes of Newborn Human Lung Cells. Genes (Basel) 15. Genes
- Minzhe Guo, Michael P. Morley, Cheng Jiang, Yixin Wu, Guangyuan Li, Yina Du, Shuyang Zhao, Andrew Wagner, Adnan Cihan Cakar, Michal Kouril, Kang Jin, Nathan Gaddis, Joseph A. Kitzmiller, Kathleen Stewart, Maria C. Basil, Susan M. Lin, Yun Ying, Apoorva Babu, Kathryn A. Wikenheiser-Brokamp, Kyu Shik Mun, Anjaparavanda P. Naren, Geremy Clair, Joshua N. Adkins, Gloria S. Pryhuber, Ravi S. Misra, Bruce J. Aronow, Timothy L. Tickle, Nathan Salomonis, Xin Sun, Edward E. Morrisey, Jeffrey A. Whitsett, NHLBI LungMAP Consortium, Yan Xu Guided construction of single cell reference for human and mouse lung. Nature Communications 14, 1-20, 2023. Nature Communications
- Minzhe Guo, Kathryn A. Wikenheiser-Brokamp, Joseph A. Kitzmiller, Cheng Jiang, Guolun Wang, Allen Wang, Sebastian Preissl, Xiaomeng Hou, Justin Buchanan, Justyna A. Karolak, Yifei Miao, David B. Frank, William J. Zacharias, Xin Sun, Yan Xu, Mingxia Gu, Pawel Stankiewicz, Vladimir V. Kalinichenko, Jennifer A. Wambach, Jeffery A. Whitsett Single cell multiomics identifies cells and genetic networks underlying alveolar capillary dysplasia. Am J Respir Crit Care Med 2023. AJRCMB
- Stevens J, Steinmeyer S, Bonfield M, Peterson L, Wang T, Gray J, Lewkowich I, Xu Y, Du Y, Guo M, Wynn JL, Zacharias W, Salomonis N, Miller L, Chougnet C, O'Connor DH, Deshmukh H. The balance between protective and pathogenic immune responses to pneumonia in the neonatal lung is enforced by gut microbiota. Sci Transl Med 2022; 14: eabl3981. PubMed bioRxiv
- Du Y, Guo M, Wu Y, Wagner A, Perl AK, Wikenheiser-Brokamp K, Yu J, Gupta N, Kopras E, Krymskaya V, Obraztsova K, Tang Y, Kwiatkowski D, Henske EP, McCormack F, Xu Y. Lymphangioleiomyomatosis (LAM) Cell Atlas. Thorax 2022: thoraxjnl-2022-218772
- Guo M, Morley MP, Wu Y, Du Y, Zhao S, Wagner A, Kouril M, Jin K, Gaddis N, Kitzmiller JA, Stewart K, Basil MC, Lin SM, Ying Y, Babu A, Wikenheiser-Brokamp KA, Mun KS, Naren AP, Lin S, Clair G, Adkins JN, Pryhuber GS, Misra RS, Aronow BJ, Tickle TL, Salomonis N, Sun X, Morrisey EE, Whitsett JA, Xu Y, Consortium NL. Guided construction of single cell reference for human and mouse lung. bioRxiv 2022: 2022.2005.2018.491687
- Toth A, Steinmeyer S, Kannan P, Gray J, Jackson CM, Mukherjee S, Demmert M, Sheak JR, Benson D, Kitzmiller J, Wayman JA, Presicce P, Cates C, Rubin R, Chetal K, Du Y, Miao Y, Gu M, Guo M, Kalinichenko VV, Kallapur SG, Miraldi ER, Xu Y, Swarr D, Lewkowich I, Salomonis N, Miller L, Sucre JS, Whitsett JA, Chougnet CA, Jobe AH, Deshmukh H, Zacharias WJ. Inflammatory blockade prevents injury to the developing pulmonary gas exchange surface in preterm primates. Sci Transl Med 2022; 14: eabl8574. PubMed bioRxiv
- Gokey JJ, Snowball J, Green J, Waltamath M, Spinney JJ, Black KE, Hariri LP, Xu Y, Perl AK. Pretreatment of aged mice with retinoic acid supports alveolar regeneration via upregulation of reciprocal PDGFA signaling.. Thorax 2021; 76: 456-467. thoraxjnl PubMed
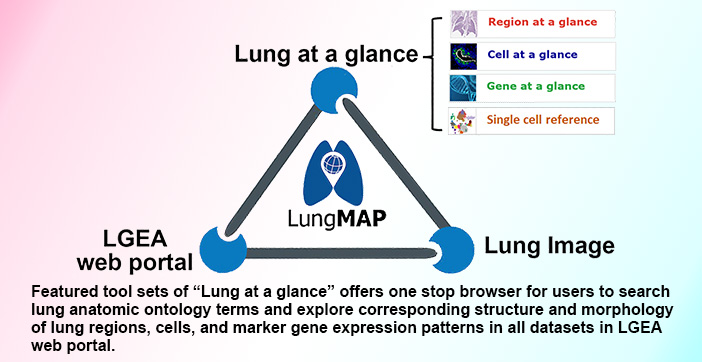
- Du Y, Ouyang W, Kitzmiller JA, Guo M, Zhao S, Jeffrey A Whitsett JA, Xu Y. Lung Gene Expression Analysis (LGEA) Web Portal v3: Lung at a Glance. AJRCMB. 2021. AJRCMB PubMed bioRxiv
- Wen B, Li E, Ustiyan V, Wang G, Guo M, Na CL, Kalin GT, Galvan V, Xu Y, Weaver TE, Kalin TV, Whitsett JA, Kalinichenko VV. In Vivo Generation of Lung and Thyroid Tissues from Embryonic Stem Cells Using Blastocyst Complementation. Am J Respir Crit Care Med 2021; 203: 471-483. AJRCMB PubMed
- Wang G, Wen B, Ren X, Li E, Zhang Y, Guo M, Xu Y, Whitsett JA, Kalin TV, Kalinichenko VV. Generation of Pulmonary Endothelial Progenitor Cells for Cell-based Therapy Using Interspecies Mouse-Rat Chimeras. Am J Respir Crit Care Med 2021; 204: 326-338. PubMed
- Sitaraman S, Martin EP, Na CL, Zhao S, Green J, Deshmukh H, Perl AT, Bridges JP, Xu Y, Weaver TE. Surfactant protein C mutation links postnatal type 2 cell dysfunction to adult disease. . JCI Insight 2021; 6. JCI PubMed
- Milewski D, Shukla S, Gryder BE, Pradhan A, Donovan J, Sudha P, Vallabh S, Pyros A, Xu Y, Barski A, Szabo S, Turpin B, Pressey JG, Millay DP, Khan J, Kalinichenko VV, Kalin TV. FOXF1 is required for the oncogenic properties of PAX3-FOXO1 in rhabdomyosarcoma. Oncogene 2021; 40: 2182-2199. Nature PubMed
- Gokey JJ, Snowball J, Sridharan A, Sudha P, Kitzmiller JA, Xu Y, Whitsett JA. YAP regulates alveolar epithelial cell differentiation and AGER via NFIB/KLF5/NKX2-1. iScience 2021; 24: 102967. PubMed
- Sun X, Perl AK, Li R, Bell SM, Sajti E, Kalinichenko VV, Kalin TV, Misra RS, Deshmukh H, Clair G, Kyle J, Crotty Alexander LE, Masso-Silva JA, Kitzmiller JA, Wikenheiser-Brokamp KA, Deutsch G, Guo M, Du Y, Morley MP, Valdez MJ, Yu HV, Jin K, Bardes EE, Zepp JA, Neithamer T, Basil MC, Zacharias WJ, Verheyden J, Young R, Bandyopadhyay G, Lin S, Ansong C, Adkins J, Salomonis N, Aronow BJ, Xu Y, Pryhuber G, Whitsett J, Morrisey EE, Consortium NL. A census of the lung: CellCards from LungMAP. Dev Cell 2022; 57: 112-145 e112. PubMed
- Bottasso-Arias N, Leesman L, Burra K, Snowball J, Shah R, Mohanakrishnan M, Xu Y, Sinner D. BMP4 and Wnt signaling interact to promote mouse tracheal mesenchyme morphogenesis. Am J Physiol Lung Cell Mol Physiol 2022; 322: L224-L242. AJPLCMP PubMed
Cite LungGENS, LGEA web portal and Lung at a glance

2. Du Y, Kitzmiller JA, Sridharan A, Perl AK, Bridges JP, Misra RS, Pryhuber GS, Mariani TJ, Bhattacharya S, Guo M, Potter SS, Dexheimer P, Aronow B, Jobe AH, Whitsett JA, Xu Y. Lung Gene Expression Analysis (LGEA): an integrative web portal for comprehensive gene expression data analysis in lung development. Thorax 2017;72(5):481-84.
3. Du Y, Ouyang W, Kitzmiller JA, Guo M, Zhao S, Whitsett JA, Xu Y. Lung Gene Expression Analysis Web Portal Version 3: Lung-at-a-Glance. Am J Respir Cell Mol Biol. 2021 Jan;64(1):146-149.
LungGENS
LungSortedCells
LungDTC
LungDisease
LungEpigenetics
LungImage
LungProteomics
LGEA-ToolBox